Latest posts:
02 Nov 2020
Europe PMC Text-annotator is now open source
Introducing Text-annotator
Europe PMC has open-sourced Text-annotator, a JavaScript library to locate and annotate plain text in HTML. The annotation process includes:
- Search: Search for a piece of plain text in the HTML; on finding it, store its location identified by an index and then return the index for later annotation.
28 Jan 2020
New release: Europe PMC Web Services 6.3
Today Europe PMC released Web Services version 6.3, updating its SOAP and RESTful APIs. Programmatic users can now find more fields available in the core response of these services
Multiple author affiliations
In the previous version, author affiliations were a single value. The new version 6.3 includes a list of multiple affiliations.
28 Jan 2020
Europe PMC API use cases page
Connecting the programmatic user community
Europe PMC makes its open access content and metadata available for building new applications. With a developer tab specially designed for programmatic users and with detailed documentation, the aim is to make it easy to integrate Europe PMC APIs into new services and thereby improve life science research.
16 Oct 2019
PDBe integrates Europe PMC APIs
PDBe exposes literature metadata by integrating Europe PMC REST APIs
PDBe, a member of the Worldwide Protein Bank, is a European resource that maintains a free and publicly available archive of macromolecular structures. The public can easily find information on protein structure and metadata associated with protein structure. PDBe also exposes enriched metadata from other sources such as publications and citations, which are retrieved through Europe PMC APIs.
20 Aug 2019
Europe PMC project for eLife Innovation Sprint
How to find the perfect preprint
The eLife Innovation Sprint is a yearly collaborative hackathon for developers, designers, researchers, technologists, science communicators, and everyone enthusiastic about open science. The premise of the Sprint is simple – the current science publishing system is slow, inefficient and insanely expensive. What we need are open science ideas that could be turned into prototypes to address the challenges we face in science publishing. All Sprint outputs have to be openly available, use open-source licenses for code and software, and permissive licenses (such as CC-BY) for other content.
This year the Europe PMC team will participate in the Sprint with a proposal to improve the discoverability of relevant scientific preprints. Dayane Araujo (Technical Outreach Officer) and Michael Parkin (Data Scientist) will be working on a tool to sort through ~80,000 life science preprints, and they need your help.
04 Jul 2018
A perfect match: locating plain text in HTML pages
SciLite is a Europe PMC tool that allows biological terms or relations, such as diseases, chemicals or protein interactions, to be highlighted for readers on abstracts and full text articles. These terms are identified as annotations by text mining algorithms, developed by a variety of text mining groups.
The main challenge for the SciLite tool is locating plain text annotations in HTML pages. The challenges derive from the nature of HTML pages. Below is a list of the major challenges we faced and the solutions adopted to mitigate them.
- The pages contain HTML tags, obviously. For example, visit this article, and click on the “Gene Function” checkbox, on the right-hand side of the page, to see the sentence highlighted.
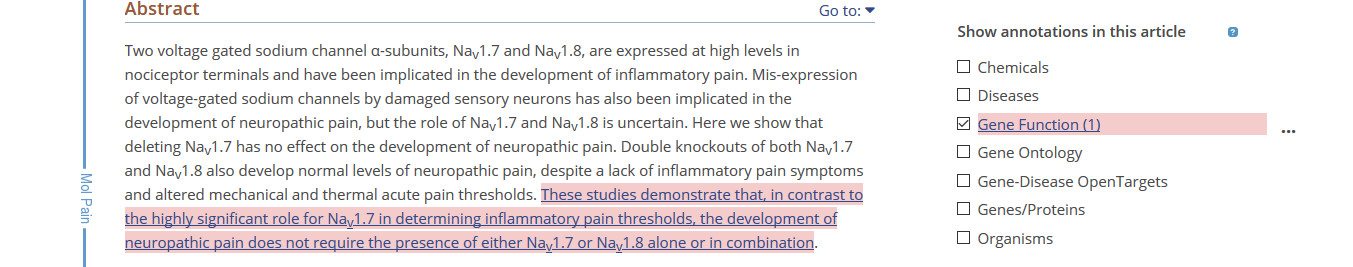 Figure 1: Annotation containing HTML tags
Figure 1: Annotation containing HTML tags
04 Jun 2018
Integrating Literature and Data
Data is at the heart of research. Scientific papers describe how data has been obtained, analysed, and what conclusions have been drawn. But it is the data that comprises the essential evidence, which confirms or disproves the original hypothesis. In the life sciences it is essential to look at scientific literature in the context of other publications, the data it builds on and other data linked to the publication. At Europe PMC we have developed a number of features to support data discovery and reuse.
As one of the ELIXIR Core Resources, Europe PMC benefits from excellent links to essential research data hubs located at EMBL-EBI. This helps us interweave publications and data, enriching the graph of research objects, and help researchers discover linked and related data.
The literature-data links come in different forms and shapes. An article might be citing a DOI for a dataset in a repository, or describe a protein structure cited as an accession number for PDBe database. An publication itself might be cited by a database, such as Flybase or even a Wikipedia article. Europe PMC obtains such literature-data links in three ways:
08 Mar 2018
The Importance of Software Testing in DevOps
Software testing is the process of identifying the correctness and quality of a software program. In other words, testing is executing a system or application in order to find software bugs, defects, errors or unexpected behavior.
Software testing is necessary because we all make mistakes. Some of those mistakes are minor, but others can be expensive or dangerous. Especially while practicing continuous integration, continuous delivery, or continuous deployment, we need to test anything and everything we produce, because things can always go wrong.
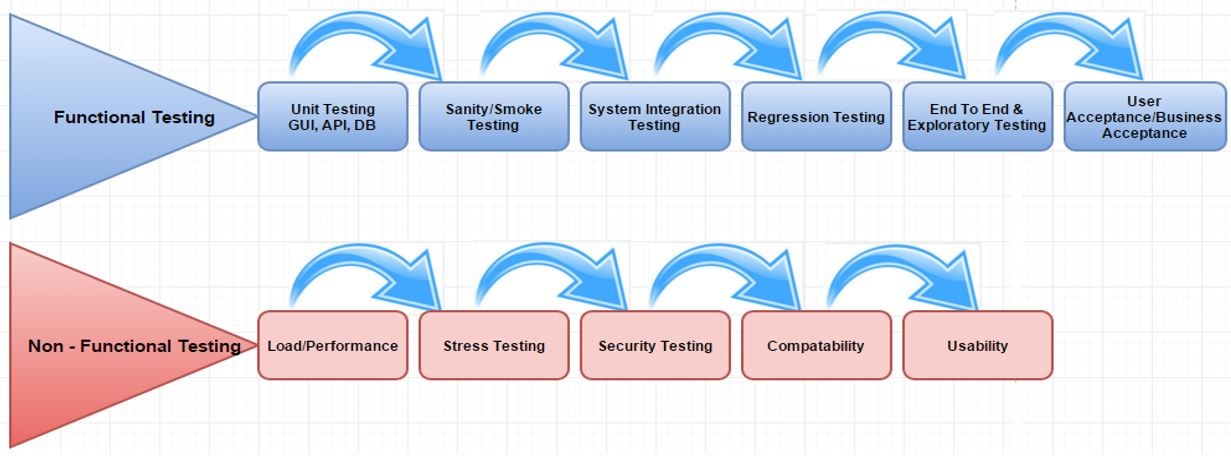
Testing is mainly classified into two types, Functional Testing and Non-functional Testing.
29 Jan 2018
Behavior-Driven Development in Bioinformatics
What is BDD?
Behavior-Driven Development (BDD) is a set of Software Engineering practices designed to help teams deliver more valuable and higher quality software features.
It adopts general techniques and principles of Test Driven Development (TDD) with ideas from Domain-driven Design (DDD). BDD incrementally builds functionality guided by expected behavior.
A simple BDD scenario / requirement is as follows:
@TC01_EPMC_SearchTest
Scenario: Specific Search by Keyword
Given I am researcher
When I open the 'Europe PMC' Website
And Enter the keyword "Glycosyl transferases" on the Query field
And Click on the Search button
Then I should be able to see the matching results on the Search Result page27 Nov 2017
Swagger documentation customisation
Swagger is a popular software framework that helps developers build RESTful Web services through their entire lifecycle, from design and documentation, to test and deployment. This post focuses on how to incorporate the API documentation generated through Swagger inside an HTML page hosted from another web application.
One of the main features of Swagger is producing interactive documentation for a RESTful API. Swagger can be used in conjunction with a multitude of different languages and frameworks. It will always produce two different outputs inside the same web application hosting the API:
- A default HTML page having a standard Swagger style. (Europe PMC Annotations API Swagger standard html page)
- A JSON file that will contain the description of the generated documentation (Europe PMC Annotations API documentation descriptor)
 Europe PMC Tech Blog
Europe PMC Tech Blog